I am currently playing with MACH and SNPMStat to impute marker based on public datasets. I do not have any opinion about the usefulness of this approach – largely due to the fact that ~10% of my allele coding is different to the public hapmap data.
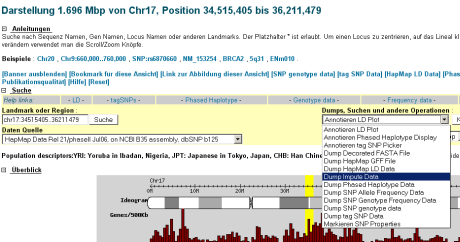
BTW the hapmap people have been very helpful in explaining to me, how imputing can be done by a web service. For that purpose llocate your region of interest then select in the dropdown “Download Impute Data” from the ‘Reports & Analysis’ section. The configuration page lets you upload a .ped and .dat files, yea, yea.
CC-BY-NC Science Surf , accessed 25.05.2026