Unfortunately with the decline of genomic research, there are some “hobbyist” researchers coming up with their own agenda. Three recent investigations lay out just how bad things have gotten – and how difficult it is to stop the damage once it starts.
The New York Times revealed that fringe researchers systematically deceived the NIH to gain access to genetic and brain-scan data from over 20,000 American children enrolled in the Adolescent Brain Cognitive Development Study. Using deliberately misleading applications, they extracted the data, shared it with unauthorized collaborators, and produced at least 16 papers purporting to rank racial groups by IQ. The papers have since been amplified millions of times on social media, cited by AI chatbots, and used as ammunition by white nationalists. The NIH eventually suspended the lead researcher and he was fired – yet his collaborators apparently retained copies of the data and kept publishing.
The Guardian then showed the problem is not limited to bad-faith actors inside the system. UK Biobank, which holds health records on 500,000 British volunteers, found that well-meaning researchers had accidentally posted sensitive datasets to GitHub dozens of times. One exposed file contained hospital diagnoses and birth details for 413,000 participants – enough to re-identify individuals with just a date of birth and one known medical procedure.
Harvard geneticist Sasha Gusev ties both stories together in a sharp Substack essay. His core point is simple: these systems were designed assuming everyone plays by the rules. They were not built for people who lie on their access applications, quietly pass data to friends, or treat a children’s health study as raw material for race propaganda. When something goes wrong, the response tends to be more paperwork, more committees, more strongly worded statements — none of which does much against someone who was never going to read them anyway. Gusev’s actual prescription: stick to what participants consented to, ban anyone who leaks data permanently, and when bad science appears, criticize it loudly in plain language where people can actually read it – not in a journal response that ten specialists will see.
The people who donated their data, often hoping to help find cures for cancer or diabetes, deserve at least that much.





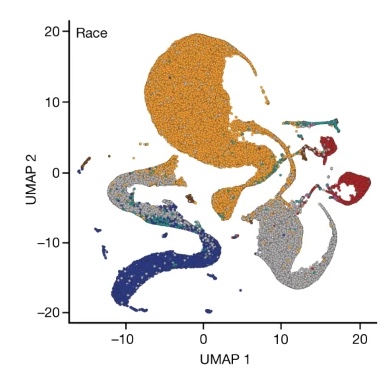 “Genetic drivers of heterogeneity” is the new description for the failed genetic concept of “reverse genetic engineering” – where we see now only sand running through the fingers.
“Genetic drivers of heterogeneity” is the new description for the failed genetic concept of “reverse genetic engineering” – where we see now only sand running through the fingers.