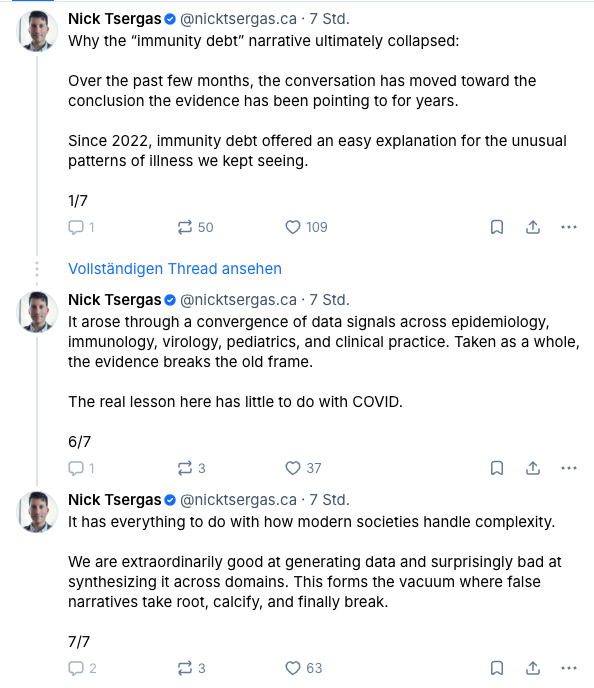
Hopefully the thread here on “farming and allergy” is now coming to an end. I tried to refute it for many years – last attempt in 2020
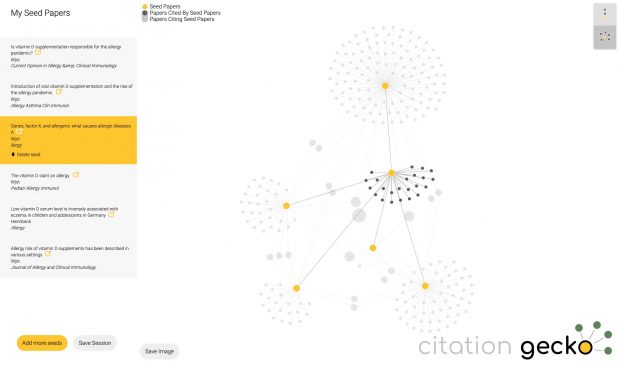
the main objection against the farming hypothesis is the interpretation of a negative statistical association as a “protective” effect. Only after thorough exclusion of alternative explanations, this interpretation may be justified.
but unfortunately missed another paper from the literature that had already been published in 2011, sorry for that.
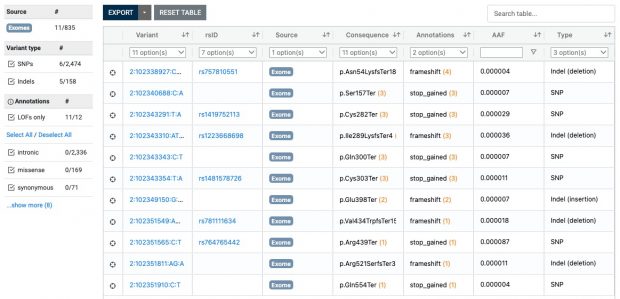
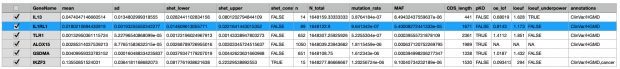
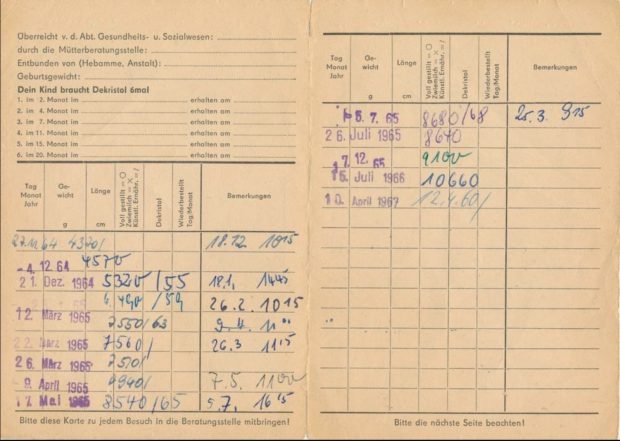
With environmental exposures aside, at least 2 profound differences between farming and nonfarming families could be a threat to the validity of such investigations. One of these is the long-term selection of traits and genes in favor of the demanding living conditions of farmers because the agricultural lifestyle is often handed down within families. Referring to the “healthy worker effect,” we could expect certain disadvantages to be underrepresented in a farming population, giving rise to the concept of a “healthy farmer effect.”
Also the second point of Grabenhenrich has never been assessed
The other area expected to be different when comparing farming and nonfarming families constitutes behavioral patterns, lifestyle, and knowledge. For example, “soft factors,” such as health care use, symptom perception, and labeling, as well as access to and interest in health-related information, are most likely to be distributed dissimilarly. This disparity, in turn, could severely affect the response pattern in studies basing their case definition mainly on questionnaires. A temporal shift of these soft factors toward increased cautiousness and awareness has been assumed to contribute to the worldwide increase in symptoms of allergic diseases and, to a lesser extent, to the increase in clinically apparent disease.10, 11 Why should such a change be uniform in all parts of a population? Particular subgroups might be susceptible to catch up more rapidly. This phenomenon could be studied by identifying outliers in otherwise homogenous populations.
in the hope that journalists and politicians will never find it?
Is this “pioneering epidemiology” as judged by MP Dr. Söder?
Promoting a stolen idea (the original was published in German only) where the source was never cited?
Is it any good science ignoring all objections?
True heroes in epidemiology and public health are Semmelweis, Snow, von Pettenkofer, Doll & Hill, Hesse & Rehn but not a MD who can not even explain an Odds Ratio…
CC-BY-NC Science Surf , accessed 13.05.2026