Sowohl das Deutsche Ärzteblatt als auch die Münchner Medizinische Wochenschrift veröffentlichten nun in kurzer Folge methodisch fragwürdige Beiträge zur Vitamin-D-Forschung. Und da sie eine enorme Reichweite in der ärztlichen Fortbildung haben, landet der Vitamin-D-Hype direkt in der Praxis mit den bekannten Folgen: Überdiagnostik, unnötige Supplementierung, wir kennen die Probleme viele Jahre zB hier. Von einer Erwiderung habe ich dennoch abgesehen, da dem Erstautor der Schlusskommentar vorbehalten bleiben sollte.
Fall 1
Die MMW druckt eine “Fortbildung”, genauer einen CME-Beitrag „Wer braucht wann wie viel Vitamin D?“ eines lange pensionierten fachfremden Arztes (MMW Fortschr Med. 2025; 167 (S3): 76–82). Leider übernimmt die KI den Unsinn nun als Expertenmeinung…
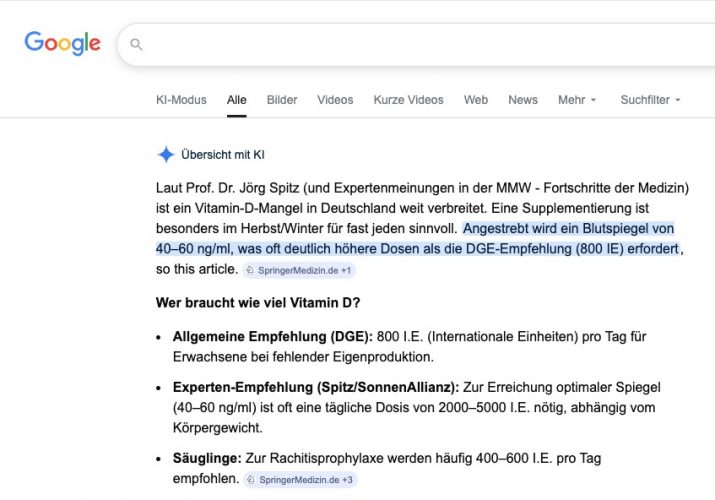
Mein Kommentar zu dem Artikel war
1. Interessenkonflikt
Der Autor gibt „keine Interessenkonflikte“ an, betreibt jedoch die Akademie für menschliche Medizin GmbH, die kommerziell Vitamin-D-bezogene Präventionsangebote vertreibt. Diese Verbindung ist gemäß DFG- und ICMJE-Regeln deklarationspflichtig. Medwatch schreibt von 225.000€ Umsatz im Jahr 2019.
2. Epidemiologische Angaben
Die Behauptung, „90 % der Bevölkerung“ hätten < 30 ng/ml Vitamin D, stützt sich auf veraltete RKI-Daten (2007–2011). Neuere Erhebungen (DEGS II, 2019) zeigen nur 30–40 % unter 50 nmol/l. → Übertriebene Darstellung eines Mangels.
3. Dosierungsempfehlung
Empfohlene 4 000–6 000 IE/Tag überschreiten den international anerkannten Upper Level von 4 000 IE/Tag (EFSA 2023). Für gesunde Erwachsene fehlt jede Evidenz.
4. Toxizität und „Coimbra-Protokoll“
Die Aussage, Werte bis 150 ng/ml seien unbedenklich, widerspricht Daten zu Hyperkalzämie und Nephrokalzinose. Das „Coimbra-Protokoll“ (≥ 100 000 IE/Tag) ist nicht evidenzbasiert und medizinisch riskant.
5. Extra-skelettale Effekte
Im Artikel werden präventive Wirkungen (Demenz, KHK, Krebs, Blutdruck, Diabetes u. a.) behauptet, obwohl große RCTs (VITAL, D2d, ViDA) keinen Nutzen zeigten. Die Darstellung ist selektiv und irreführend.
6. Schwangerschaft
Die genannte 60 %ige Reduktion von Frühgeburten entstammt keiner belastbaren Quelle; die zitierte Pilotstudie (Wagner et al. 2006) weist diese Endpunkte nicht auf.
7. COVID-19-Bezug
Die angeblich 16-fache Mortalitätssteigerung bei Vitamin-D-Mangel beruht auf einem Meinungsartikel, nicht auf einer Originalstudie. Die Darstellung ist faktisch falsch.
8. Fehlende Neutralität im CME-Kontext
Der Beitrag enthält werblich anmutende Aussagen, selektive Quellenwahl (u. a. Masterarbeit Göthel 2020) und unkritische Übernahme von Hypothesen. Damit ist die formale Neutralitätsanforderung der CME-Zertifizierung (§ 4 CME-Richtlinie BÄK) nicht erfüllt.
Der Chefredakteur Cornelius Heyer nimmt auf meine Reklamation zwar die CME Akkreditierung heraus, aber statt einer Korrektur oder Löschung des Beitrages, druckt die MMW einen Leserbrief nach dem Spitz das letzte Wort in der Diskussion hat und tatsächlich in Bhakdi Manier schreibt “Die von mir dargestellten Fakten entsprechen der aktuellen internationalen Forschung – wenn die Leitlinien das nicht tun bitte ich, dies mir nicht anzulasten”.

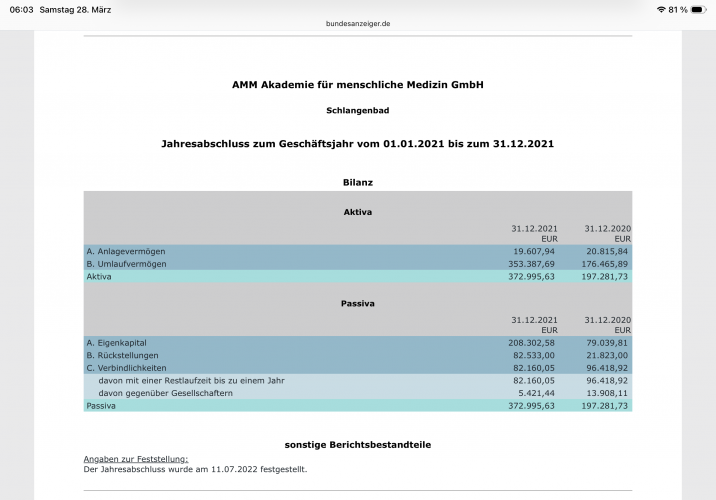
Ein beliebiger Leser der Tageszeitung ist damit nun also besser informiert als ein/e Arzt/Ärztin der die MMW abonniert hat.
Fall 2
Das Deutsche Ärzteblatt druckt ein Review eines Kongressabstracts “Personalisierte Vitamin D Supplementierung kann das Re-Infarktrisiko halbieren”. Wer nec ist weiss ich nicht, allerdings weiss ich genau, dass es wieder Märchen sind die nec hier erzählt. Der Leiter der Medizinisch-Wissenschaftlichen Redaktion des DÄ Christopher Baethge verweist mich an die journalistische Redaktion in Berlin Michael Schmedt, der allerdings auch nach Wochen immer noch nicht geantwortet hat.
New Orleans – Eine Supplementierung mit Vitamin D in individuell titrierter Dosierung kann das Risiko für einen erneuten Herzinfarkt bei vorerkrankten Personen um mehr als die Hälfte reduzieren. Das berichteten Forschende bei den Scientific Sessions 2025 der American Heart Association in New Orleans (Abstract Nr. 4382525).
[…]
Warum ist das so?
Die TARGET-D-Studie wird als Korrektur früherer negativer Vitamin-D-Studien präsentiert, mit der Behauptung, diese seien gescheitert, weil sie Vitamin D nicht „zielgerichtet“ titriert hätten. Bei genauer Betrachtung reproduziert TARGET-D jedoch genau jene strukturellen Schwächen, die die Vitamin-D-Outcome-Literatur seit Jahren geplagt haben.
Der entscheidende Befund ist eindeutig: Der vordefinierte primäre Endpunkt, schwere kardiovaskuläre Ereignisse (MACE), wurde in der Intention-to-treat-Analyse nicht signifikant reduziert. Nach elementaren CONSORT-Prinzipien müsste dieses Ergebnis die Interpretation bestimmen. Stattdessen verlagert die Studie den Fokus sofort auf ein einzelnes positives Signal in einer Unterkomponente des kombinierten Endpunkts, den Folge-Myokardinfarkt. Diese Verschiebung rettet die Studie nicht, sondern verschleiert ihr negatives Hauptergebnis. Wenn ein kombinierter Endpunkt scheitert, stellt die selektive Hervorhebung einer einzelnen Komponente keine Evidenz dar, sondern narrative Verzerrung.
Die anschließende Betonung von Per-Protocol-Analysen untergräbt die Aussagekraft weiter. Diese Analysen vergleichen nicht mehr randomisierte Gruppen, sondern Untergruppen, die nachträglich anhand des Erreichens eines willkürlich gewählten Vitamin-D-Schwellenwertes definiert werden. Damit wird nicht mehr die Wirkung einer Intervention geprüft, sondern der Gesundheitszustand einer selektierten, therapietreuen, Responder Population dem einer Restgruppe gegenübergestellt, die überproportional Nicht-Responder, Gebrechliche und Patienten mit höherer Krankheitslast enthält. Der Vitamin-D-Spiegel fungiert hier als Marker guter Gesundheit und Compliance, nicht als kausaler Faktor. Genau dieser Denkfehler hat frühere Beobachtungsstudien diskreditiert, was auch in allen Umbrella Reviews klar herauskam, aber hier innerhalb eines randomisierten Designs erneut eingeführt wird.
Das Ausmaß der postrandomisierenden Selektion macht dieses Problem unübersehbar. Ein erheblicher Teil der dem Vitamin-D-Arm zugewiesenen Patienten wird aus der Per-Protocol-Analyse ausgeschlossen, weil der Zielwert nie erreicht wurde oder nur unvollständige Nachbeobachtung vorlag. Die Konditionierung auf einen postrandomisierten Biomarker zerstört die durch Randomisierung erreichte Vergleichbarkeit der Gruppen. Die daraus resultierenden Effekte sind selektionsgetrieben und nicht kausal interpretierbar.
Auch die biologische Grundannahme der Studie ist schwach fundiert. Der gewählte Zielwert von über 40 ng/ml für 25-Hydroxyvitamin D ist nicht leitlinienbasiert, entspricht nicht der neuesten Literaturund impliziert einen Schwellen- oder Dosis-Wirkungs-Effekt, der in großen randomisierten Studien und genetischen Analysen nicht bestätigt wurde. Indem der Studienerfolg über das Erreichen dieses Zielwerts definiert wird, wird der behauptete Nutzen nicht getestet, sondern implizit vorausgesetzt.
Die Darstellung der Ergebnisse verstärkt diesen Eindruck. Formulierungen wie „klinisch relevante Risikoreduktionen“ oder „Reduktion des Myokardinfarktrisikos um mehr als die Hälfte“ suggerieren einen kausalen Effekt, den das Studiendesign und die Resultate nicht tragen. Andere Komponenten des kombinierten Endpunkts zeigen keine konsistente Verbesserung, werden jedoch deutlich weniger betont. Der Gesamteindruck ist nicht der einer neutralen Prüfung einer Hypothese, sondern der Versuch, aus einer im Kern negativen Studie ein positives Narrativ zu extrahieren.
Insgesamt liefert TARGET-D keinen belastbaren Beleg dafür, dass eine Vitamin-D-Normalisierung das kardiovaskuläre Risiko nach akutem Koronarsyndrom senkt. Der primäre Endpunkt ist negativ, die sekundären Aussagen beruhen auf selektiver Gewichtung, und die Per-Protocol-Analysen ersetzen Randomisierung durch Adhärenz- und Selektionsmechanismen. Die Studie bestätigt letztlich nur, was die Literatur seit Langem zeigt: Niedrige Vitamin-D-Spiegel korrelieren mit schlechter Gesundheit, ihre Korrektur verändert jedoch harte kardiovaskuläre Endpunkte nicht zuverlässig. Neu an TARGET-D ist nicht die Überwindung dieser Limitationen, sondern ihre methodisch aufwendigere und potenziell irreführende Verpackung.
Die Assistentin der Chefredaktion Monia Jacobs entgegnet sinngemäß am 15.4. per Email, die Studie sei auf dem renommierten AHA-Kongress vorgestellt und von der AHA selbst per Pressemitteilung kommuniziert worden – ergo nicht das Problem des Ärzteblattes. Der eigene Beitrag würde die Ergebnisse transparent darstellen, einschließlich des verfehlten primären Endpunkts. Die Umdeutung ist aber m.E. narrative Verzerrung. Journalistische Sorgfaltspflicht verlangt, genau das zu benennen – sofern man denn überhaupt verstanden hat worum es hier geht.