Response to oral vitamin D seems to be different in humans . How do we buffer (artificial) vitamin D intake?
Vitamin binding protein or group specific component GC is a good candidate. GC regulates the bioavailability of 25(OH)D3, acting as the main transporterint he blood stream from liver to kidney. As described earlier GC binds with high affinity to 25(OH)D3, leaving less than 1% of circulating 25(OH)D3 free. In contrast to 25(OH)D3, which has a half-life of several weeks, GC has a short half-life of 3 days only, suggesting that the protein and its ligand are independently regulated. Also the free binding capacity of GC is variable. In addition there are GC variants that have different binding characteristics. Depending on these isoforms, serum levels increased between 97% and 307% after receiving 600 or 4000 IU/d vitamin D3 for one year. Taken together GC is assumed to be a buffer of vitamin D effects (and side effects) whenever transport in the blood stream is being involved.
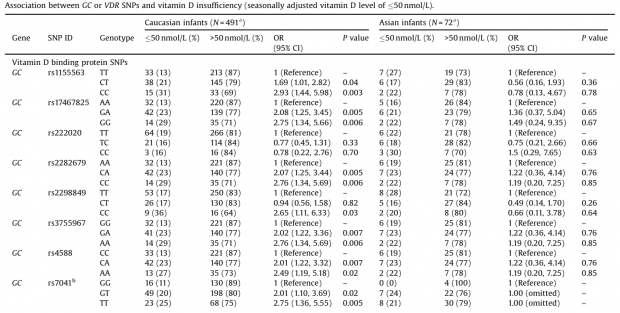
The most recent GWAS study now shows again skyrocking p-values of GC variants and serum 25(OH)D3.
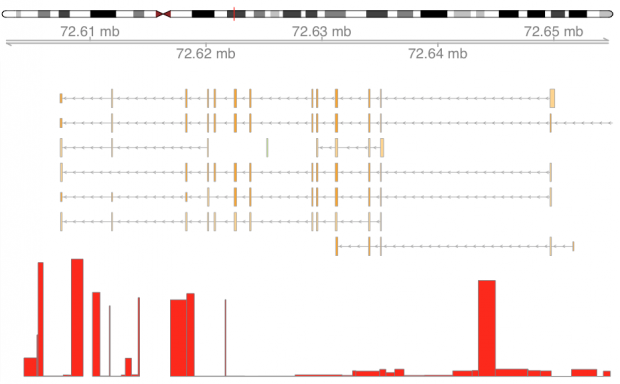
It is long known, that two missense variants of GC locate in exon 11. rs7041 encodes Asp432Glu pr D432E and rs4588 encodes Thr436Lys or T436K. These amino acid exchanges are leading to electrophoretically distinguishable proteins Gc1F/Gc1S and Gc2 respectively. We are moving the following gene plot bottom up to match the orientation.
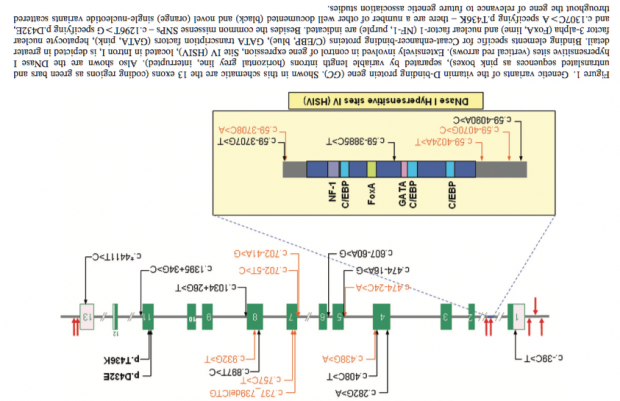
Unfortunately LD is extremely high at GC. The GWAS peaks are therefore in the first intron, at exon 11 and intron 12. Lets’ s get closer to exon 11 where the two most important SNPs reside.
Although both variants are listed at many SNP chips I can find only results for rs7041 with p=10^-222 in the new dataset.
![]()
rs7041 is listed as a A->C SNP there but according to Fu 2009 it is definitely a G->T variant. Also SNPedia has numerous articles for rs7041 being a G->T exchange, for example Suaini 2014
This is also confirmed by dbsnp. The GAT -> GAG exchange is equivalent to D -> E, so the online results report a wrong strand orientation. Unfortunately we are stuck here, as one of the main effect SNP seems to have an unclear allele assignment and the second most important SNP is missing from the meta-analysis.
What would be nice is a conditional analysis based on rs7041/rs4588 haplotypes. I predict there are further unknown functional variants in GC. Maybe in intron 1 that often contains regulatory elements at the 5′-site of the intron. As the strongest signal is in the last intron and even beyond the 3′- end, further studies of 3′-UTR would be interesting, looking for binding sites of regulatory proteins, some miRNA or AU rich elements that affect the stability or decay rate of the transcript.