Eric Topol will agree on the fact that
the practice of utilizing the DNA of an individual to predict disease has been judged to provide little to no useful information.
but they nevertheless the group tried to rescue the concept by combining clinical risk factors plus low/medium/high polygenic risk. As a pragmatic approach this may work for some diseases but it will not explain any genetic pathhway.
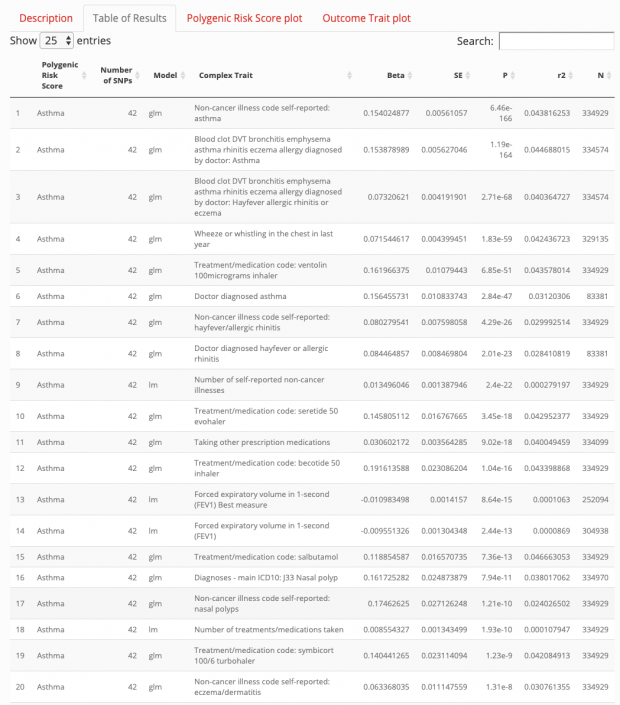
More recently Richardson et al. tried another approach using the UK Biobank data where they analysed 162 PRS and 551 heritable traits from 334,398 individuals. Using their web application for my search term “asthma” I did not find anything useful that hasn’t been known for ages.
So lets have a look at an example published last November in Cell “Screening Human Embryos for Polygenic Traits Has Limited Utility”. The outcome is sobering: If an IVF embryo would be profiled with polygenic scores for traits such as height or IQ
the top-scoring embryo is expected to be about 2.5 cm or about 2.5 IQ points above the average. The adult trait value of the top-scoring embryo would remain widely distributed.
I wouldn’t have even run this analysis for being pointless.