All posts by admin
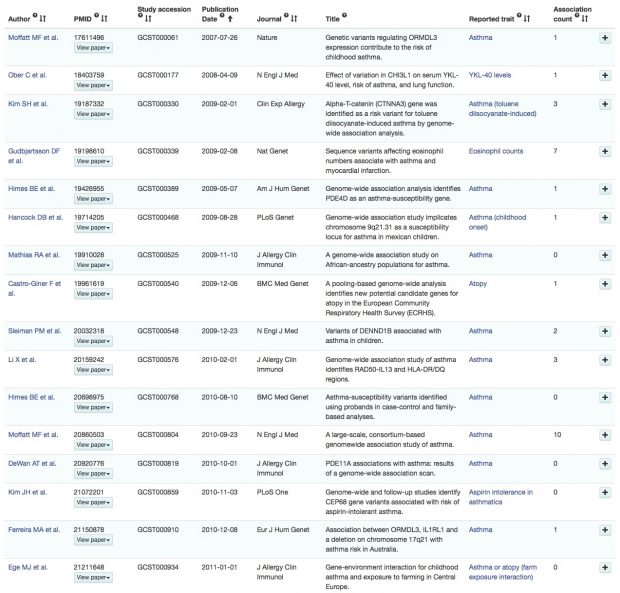
IL1RL1 / ST2 and asthma
The association of IL1RL1 and asthma is usually attributed to our 2009 nature genetics paper, sometime also to the later NEJM paper 2010. We described the association in the IL1 cluster, however, already 6 years earlier in Gohlke et al. 2004. The resolution wasn’t so good at that time as we thought more of IL1RN than IL1RL1. And even in 2004, the IL1 cluster was well known. Why is the temporal order of events so unimportant?
Success!
Beutegemeinschaften
Ulrich Dirnagl hat ein lesenswertes Stück im LJ 11/2017, S.25
… Wir arbeiten in Gruppen von im Schnitt acht Forschern (inklusive Studenten), die durchweg Arbeitsverträge über wenige Jahre haben – und mit Fördergeldern, die höchstens für drei Jahre gesichert sind. Die Forschungsstrategien und Ergebnisse werden bis zur endgültigen Publikation unter Verschluss gehalten, man könnte ja gescoopt werden.
Data Sharing? Um Himmels willen. Hab’ ich ja nichts davon, könnte im Zweifelsfall sogar schaden. Wie verrückt aber ist die Frage, ob sich die grundlegenden Fragen der Biowissenschaften und der Medizin nicht besser in multinationalen, koordinierten sowie ausreichend und langfristig alimentierten Kooperationsprojek-ten aufklären ließen? Sollte man das wenigstens nicht mal ausprobieren? Continue reading Beutegemeinschaften
Why replication is important
A nice explanation at honestuniverse.com
One of the most important principles of the scientific method is reproducibility. A valid result should be able to be replicated independently, whereas an invalid result (originally achieved due to some error or perhaps just chance) will not be able to be consistently reproduced […] The reason why this is the case has to do with researcher degrees of freedom. In an original experiment, the experimenters have the freedom to make certain choices. Some choices may have been made beforehand, whereas others are made after the study has been started. These choices may not all be made consciously, or they may be made consciously with only unconscious bias, but the fact that the choices are made at all after the experiment has begun affects the reliability of the results.
which basically means that the kind of replication nature genetics demands for an association, is not a replication at all, but some extended or repeated study.
How to merge local and remote data in the browser
In a first step reconfigure your browser
Safari
Enable the develop menu using the preferences panel, under Advanced -> “Show develop menu in menu bar”.
Then from the safari “Develop” menu, select “Disable local file restrictions”.
Chrome
Close all running Chrome instances first. Then start the Chrome executable with a command line flag:
chrome –allow-file-access-from-files
On Mac OSX terminal you can do this with
open /Applications/Google\ Chrome.app –args –allow-file-access-from-files
Firefox
In the address bar, type about:config Find the security.fileuri.strict_origin_policy parameter and set it to false
Here is the script, loosedly based on https://www.html5rocks.com/en/tutorials/file/dndfiles and https://stackoverflow.com/questions/7346563/loading-local-json-file that loads json1 from the server, json2 from the local harddrive and merges both by id
<html>
<input type="file" id="files" name="files[]" />
<output id="list"></output>
</html>
<script src="jquery-1.11.1.min.js"></script>
<script>
// read local comma delimited file with id and name
function readTextFile(file) {
var rawFile = new XMLHttpRequest();
rawFile.open("GET", file, false);
rawFile.onreadystatechange = function () {
if(rawFile.readyState === 4) {
if(rawFile.status === 200 || rawFile.status == 0) {
rider = rawFile.responseText.split("\n");
return rider;
}
}
}
rawFile.send(null);
}
// read server data in json format
var url = 'merge.json';
var json2;
$.ajax ({
url: url,
async: false,
dataType: "json",
cache: false,
success: function(data) {
json2 = data;
},
error: function(err) {
console.log(JSON.stringify(err))
}
});
// function to join by id
function joinObjects() {
var idMap = {};
for(var i = 0; i < arguments.length; i++) {
for(var j = 0; j < arguments[i].length; j++) {
var currentID = arguments[i][j]['id'];
if(!idMap[currentID]) {
idMap[currentID] = {};
}
for(key in arguments[i][j]) {
idMap[currentID][key] = arguments[i][j][key];
}
}
}
var newArray = [];
for(property in idMap) {
newArray.push(idMap[property]);
}
return newArray;
}
// selection of local fiel starts it all
function handleFileSelect(evt) {
var files = evt.target.files; // FileList object
// files is a FileList of File objects. List some properties.
var output = [];
for (var i = 0, f; f = files[i]; i++) {
readTextFile( "file:///Users/wjst/Desktop/"+escape(f.name) );
for(var i = 0; i < rider.length; i++) {
rider[i] = rider[i].split(",");
}
json1 = rider.map(function(x) {
return {
"id": Number(x[0]),
"name": x[1]
}
})
console.log(json1);
console.log(json2);
console.log( JSON.stringify( joinObjects(json1, json2) ));
}
}
document.getElementById('files').addEventListener('change', handleFileSelect, false);
</script>
Wissenschaftsrat zu Peer Review
wissenschaftsrat.de schreibt
Das Begutachtungswesen (peer review) steht nach Ansicht des Wissenschaftsrates unter zunehmendem Druck. Sowohl die Erwartungen an die Leistungen von Gutachterinnen und Gutachtern als auch die Nachfrage nach Begutachtungen seien aus verschiedenen Gründen deutlich gewachsen, ohne dass die Gruppe der Gutachtenden systematisch vergrößert und Wissen über Begutachtungen strukturierter vermittelt wurden.
Positionspapier 23.10.2017
Treg and transplant rejection
Yesterday evening I heard an interesting talk here about lung transplant rejection where one of the best prognostic marker
was the (pre transplantation) Treg count.
I wonder if there are some preexisting patient conditions, like allergy, that are responsible for this phenomenon?
At least it is already known since 2008 that Treg function is important. Current Treg therapies are exploiting this. Another allergy hallmark the IL33/ST2 axis is actively tested at the moment in transplant rejection.
In contrast transplant acquired allergy is something different – a side effect of tacrolimus? Or any other ingredient of the cocktail of steroids, mycophenolate, and IL-2 receptor inhibitors such as basiliximab?
Exports OSX ratings to XMP ratings
for FILE in *.jpg
do
FN=$(echo $FILE | sed -e "s/jpg/xmp/")
RATE=$(mdls "$FILE" | awk '/kMDItemStarRating/' | awk '{print $NF}' )
OR=$(mdls "$FILE" | awk '/kMDItemOrientation/' | awk '{print $NF}' )
exiftool "$FILE" -xmp:rating=$RATE -n -xmp:orientation=$OR -o "$FN"
done
Keep your Excelsheet updated ;-) or how to bind Kendo UI for jquery on a Synology to websocket
This is not really about Excel but about Kendo Grid’s ability to update grid data in the browser. This is already my second attempt as a Mosquitto based MQTT solution did not work properly.
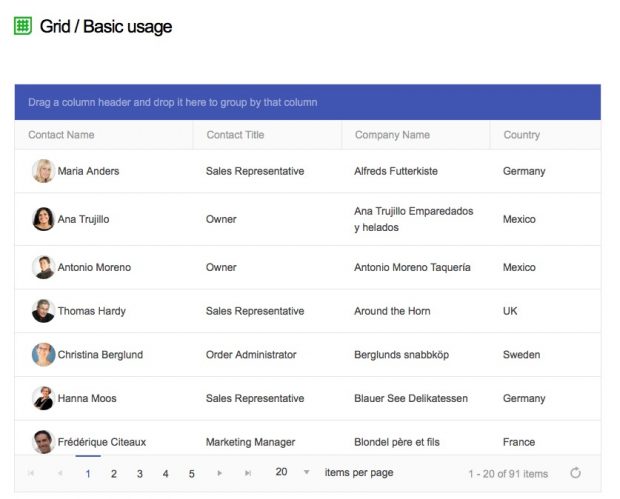
Installing node.js on the Synology is straightforward – just use the package manager. We also need some local packages. Change into the target directory and type
npm install express npm install typescript npm install ws
Copy also app.js from https://github.com/telerik/kendoui-ws-demo into that directory. Unfortunately app.js breaks in line 5 (we need another port as 5000 is already occupied in the Synology). It also crashes in line 6, as the node.js V4 version needs full extension. So use the following lines
var WebSocketServer = require("ws").Server;
var http = require("http");
var express = require("express");
var app = express();
var port = process.env.PORT || 5050;
var products = require("./products.js");
Then start the server by
/usr/local/bin/node /volume1/web/kendoui-ws-demo-master/app.js &
Get the client script in kendoui.for.jquery.2017.3.1018.trial/examples/grid/web-socket.html and change the host. Take care to use ws (Websocket), not wss (Websocket secure) unless you know what you are doing.
//var host = "wss://kendoui-ws-demo.herokuapp.com"; var host = "ws://myserver:5050";
That’s it.
First allergy cell found
Allergen-specific type 2 helper T (TH2) cells play a central role in initiating and orchestrating the allergic and asth-matic inflammatory response pathways. One major factor limiting the use of such atopic disease-causing T cells as both therapeutic targets and clinically useful biomarkers is the lack of an accepted methodology to identify and differentiate these cells from overall nonpathogenic TH2 cell types. We have described a subset of human memory TH2 cells confined to atopic individuals that includes all allergen-specific TH2 cells. These cells are terminally differ-entiated CD4+ T cells (CD27− and CD45RB−) characterized by coexpression of CRTH2, CD49d, and CD161 and exhibit numerous functional attributes distinct from conventional TH2 cells. Hence, we have denoted these cells with this stable allergic disease-related phenotype as the TH2A cell subset.
Sounds promising…
Hygiene hypothesis should be death but is still alive
There is a new paper today “The Hygiene Hypothesis in the Age of the Microbiome“. Fortunately there is a strong paywall…
An impressive illustration of the hygiene hypothesis was found in the consistent farm effect on asthma and allergies, which has partly been attributed to immunomodulatory properties of endotoxin as emitted by livestock.
which is not aware of “Time to abandon the hygiene hypothesis: new perspectives on allergic disease, the human microbiome, infectious disease prevention and the role of targeted hygiene”
Although evidence supports the concept of immune regulation driven by microbe–host interactions, the term ‘hygiene hypothesis’ is a misleading misnomer.
see also “News Feature: Cleaning up the hygiene hypothesis”
Still, the catchy hygiene hypothesis continues to be widely embraced by the public, the media, and even scientists: Uses of the term in the scientific literature rose threefold over the past 10 years compared with the decade prior, according to a search on Thomson Reuters Web of Science. “In science, when something has been propagated for so long, it can be hard to change,” says Marsha Wills-Karp, chair of environmental health and engineering at Johns Hopkins Bloomberg School of Public Health.
Timeline asthma / allergy genetics
First Genetic Study
1923 AF. Coca.J Immunol 8:163-171
First Linkage Study (AB0 blood group)
1936. I. Zieve, Ann Hum Gen 7:4074, 163-178
First Molecular Study (HLA)
1973 D. Marsh, Science 179:4074, 691-3
Second Linkage Study (SLC14A1)
1985, H. Eiberg, Cytogenet Cell Genet 40: 622 -journal discontinued, no online archive-
Third Linkage Study (11q31) – questionable results –
1989 W. Cookson, Lancet 1:8650, 1292-5
First Modern Family Study
1992 S. Dold, Arch Dis Child 67:8, 1018-22
First gene (Fc∈RI-β)
1993 A.J.Sandford, The Lancet 341:8, 332-334
First true gene (IL4)
1994 D. Marsh, Science 264:5162, 1152-6
First Genomewide Linkage Scan
1996 S. Daniels, Nature 383:6597, 247-50
Transatlantic Airway Conference Key Biscayne FL
1997 N. Zamel (only reported later by Thompson 2003 and Duffy 2019)
Second Genomewide Linkage Scan
1997 CSGA Nature Genetics 15:389-392
Third Genomewide Linkage Scan
1999 M. Wjst, Genomics 58:1-8
First GWAS
2007 M. Moffatt, Nat Genet 15:389-392
Second GWAS
2008 C. Ober, NEJM 358: 1682-91
Phantom reference
There is an interesting story at harzing.com: A conference template with a dummy reference, that was supposed to be edited by abstract authors, was finally cited by 400 authors :) I find even more citations at Google Scholar, 703 to be exact. Low quality, missing control, careless editing, and fake journals, creepy…
Open letter to the Sente developer
Dear Michael J. Cinkosky,
this is an open letter regarding Sente academic library software.
Sente is used by tens of thousands of researchers in all academic fields at universities and research organizations around the world
I am using Sente nearly every day for many years to collect and annotate the literature.
It is useful to retrieve papers and insert references in my own papers.
It seems that Sente is not more supported.
A while ago, I could build up a bibliography only in exports from my text processor.
Then online sync stopped.
Most recently also Google Scholar reference recognition ended.
It is now a useless software. I paid for it, I used it, which was a fair deal.
But now am now lost, if Sente just drowns in the internet.
Could you sell it to a company that continues support?
Your customer base would really appreciate it.
Or write a final countdown Endnote export script that preserves PDF links.
Or just give us the code at Github, we will continue to develop it.
We all appreciate your work with Genbank and also with Sente.
But don’t leave us without saying good-bye.
All the best,