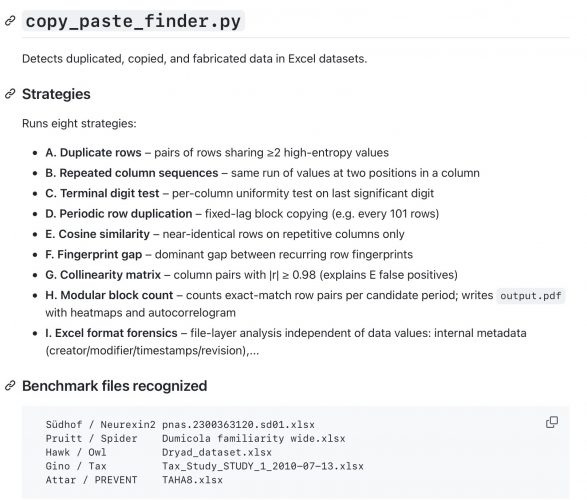
This is a quick shot of a Python based Excel data analysis tool that is based on some earlier findings described at here
Just head over to Github for the most recent version.
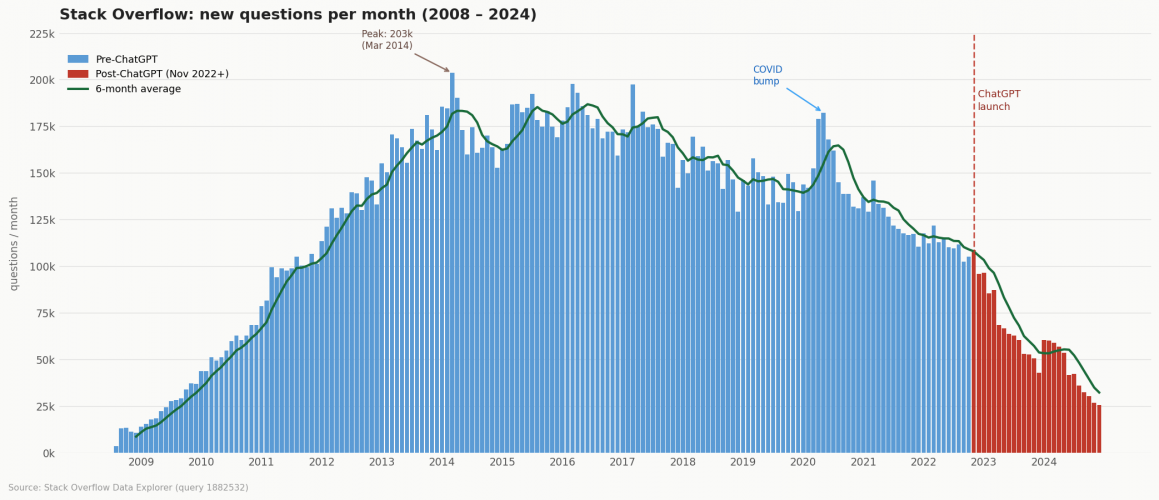
Coding with AI has a nice chart, that I am redrawing here
so it is time to say Good-Bye now after 14 years
and sticking to the new 10 commandments by Russell Poldrack
Gather Domain Knowledge Before Implementation
Distinguish Problem Framing from Coding
Choose Appropriate AI Interaction Models
Start by Thinking Through a Potential Solution
Manage Context Strategically
Implement Test-Driven Development with AI
Leverage AI for Test Planning and Refinement
Monitor Progress and Know When to Restart
Critically Review Generated Code
Refine Code Incrementally with Focused Objectives
There is an interesting observation by Nick Brown over at Pubpeer who analysed a clinical dataset (see also my comment atthe BMJ)
…there is a curious repeating pattern of records in the dataset. Specifically, every 101 records, in almost every case the following variables are identical: WBC, Hb, Plt, BUN, Cr, Na, BS, TOTALCHO, LDL, HDL, TG, PT, INR, PTT
which is remarkable detective work. By plotting the full dataset as a heatmap of z scores, I can confirm his observation of clusters after sorting for modulo 101 bin.
How could we have found the repetitive values without knowing the period length? Is there any formal, data-agnostic detection method?
If we even don’t know the initial sorting variable, it may makes sense to look primarily for monotonic and nearly unique variables, i.e. that are plausible ordering variables. Clearly, that’s obs_id in the BMJ dataset.
Let us first collapse all continuous variables of a row into a string forming a fingerprint. Then we compute pairwise correlations (or Euclidean distances in this case) of all fingerprints. If a dataset contains many identical or near-identical rows, we will see a multimodal distribution of correlations plus an additional big spike at 1.0 for duplicated rows. This is exactly what happens here.
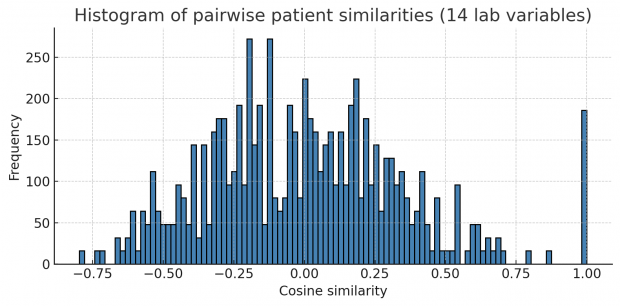
Unfortunately this works only when mainly repetitive variables are included and not too many non repetitive variables.
Next, I thought of Principal Component Analysis (PCA) as the identical blocks may create linear dependencies and the covariance matrix is becoming rank-deficient. But unfortunately results here were not very impressive – so we better stick with the cosine similarity above.
So rest assured we find an excess of identical values, but how to proceed? Duplicates spaced by a fixed lag will cause an high lag k autocorrelation in each variable. Scanning k=1…N/2 reveals spikes at the duplication lag as shown by a periodogram of row-wise similarity in the BMJ dataset.
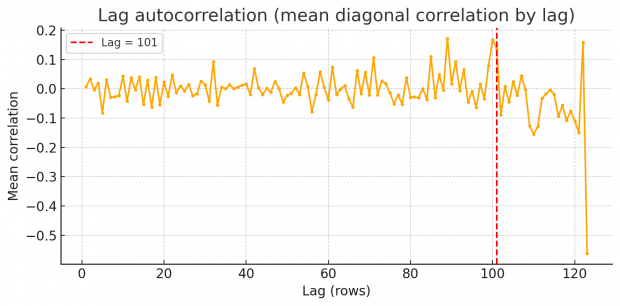
So there are peaks at around 87, 101 and 122. Unfortunately I am not an expert in time series or signal processing analysis. Can somebody else jump in here and provide some help with FFT?
There may be even an easier method, using the fingerprint-gap . For every fingerprint that occurs more than once, we sort those rows by obs_id and compute the differences of obs_id between consecutive matches. Well, this shows just one dominant gap at 101 only!
We could test also all relevant mod values, lets say between 50 and 150. For each candidate we compute the across-group variance of the standardized lab-means. The result is interesting
Modulus 52: variance = 0.084019 Modulus 87: variance = 0.138662 Modulus 101: variance = 0.789720
As a cross check let us look into white blood cell counts (WBC) and hemoglobin (Hb).
I am not sure, how to interpret this. Mod 52 may reflect shorter template fragments but did not show up in the autocorrelation test. Mod 87 has rather smooth, coherent curve and is supported by autocorrelation. Mod 101 is more noisy, but gives probably the best explanation for block copying values. Maybe the authors block copied at two occasions?
On the next day, I thought of a strategy to find the exact repetition numbers. Why not looping over mod 50 through 150 and just count the number of identical blocks? This is very informative – blocks of size 2, size 3 and 4 or greater show an exact maximum at modulus 101.
23.3.2026 Appendix
There seems many more studies out there with copy-pastein signs including a Parkinson Cell paper, a PLoS Genetics toxicology paper and a Nat Comm fish ecology study. Here is the Github link to the implementation by Markus Eglund
Hopefully I get the pipeline right by summarizing the entropy calculation there. This is not Shannon entropy – it is a custom measure of how informationally surprising a raw number is. The logic is:
0.314 → 314, 0.500 → 5 (trailing zeros stripped), 2016 → 16 (year exception: years 1900-2030 get a capped entropy of 100).log10(value); values up to 100,000 get 5×log10 - 8; larger values get log10 + 12.logNumberCountModifier (log of the total number of numeric cells on the sheet) so large sheets don’t get disproportionately penalised.The suspicion grades are fixed thresholds on the resulting normalized score. I will add the strategy to my Python script (it is implemented here in type script) as another module and upload to Github once it has been sufficiently tested.
31.3.2026 Appendix
PREVENT-TAHA8, the starting point of this analysis, has been retracted today. I will give a presentation on the avalanche, that has been triggered by this paper, on 29–31 July 2026 in Hannover.
# v1 conda env export --from-history > environment.yml conda env create -f environment.yml # v2 conda install conda-pack conda pack -n myenv -o myenv.tar.gz # on target system mkdir -p ~/envs/myenv && tar -xzf myenv.tar.gz -C ~/envs/myenv ./bin/conda-unpack # v3 www.docker.com # v4 micromamba env export / micromamba pack
There are only a few photographs that made headlines recently. One is Man Ray’s Le Violon d’Ingres for its price tag of $12,400,000.
Or the authorship discussion around the “Napalm Girl” Phan Thị Kim Phúc.
And there is a third photograph – a snapshot from a London townhouse two decades ago – that has a similar price tag attached like Le Violon d’Ingres.

My recent paper at https://arxiv.org/abs/2507.1223 examines this infamous photograph using the latest image analysis techniques.
This study offers a forensic assessment of a widely circulated photograph featuring Prince Andrew, Virginia Giuffre, and Ghislaine Maxwell – an image that has played a pivotal role in public discourse and legal narratives. Through analysis of multiple published versions, several inconsistencies are identified, including irregularities in lighting, posture, and physical interaction, which are more consistent with digital compositing than with an unaltered snapshot. While the absence of the original negative and a verifiable audit trail precludes definitive conclusions, the technical and contextual anomalies suggest that the image may have been deliberately constructed. Nevertheless, without additional evidence, the photograph remains an unresolved but symbolically charged fragment within a complex story of abuse, memory, and contested truth.
I provide also a 3D reconstruction of the scene in the preprint although some people may find it easier to watch a video instead.
Even after completion of the analysis there are many open questions – where is the original headshot? There are numerous similar images at various image archives while I have not found any 100% original copy so far.


Even as there are now reasonable doubts on the image, Prince Andrew could have of course met Virginia Giuffre. Maybe like an artist is painting a scene from memory, this photograph could be showing a real scene although clearly not in a physical sense.
So, to repeat my last sentence in the paper – this photograph remains an unresolved but symbolically charged fragment within a complex story of abuse, memory, and contested truth.
Note added March 2, 2026
Will add an update to the preprint in the next week as there are some interesting new results from the a comparative analysis of the above images.


Here is the link to the famous landmark paper in the recent history https://arxiv.org/abs/1706.03762
Before this paper, most sequence modeling (e.g., for language) used recurrent neural networks (RNNs) or convolutional neural networks (CNNs). These had significant limitations, such as a difficulty with long-range dependencies and slow training due to sequential processing. The Transformer replaced recurrence with self-attention, enabling parallelization and faster training, while better capturing dependencies in data. So the transformer architecture became the foundation for nearly all state-of-the-art NLP models. This enabled training models with billions of parameters, which is key to achieving high performance in AI tasks.
The recent Science Advance paper by Kobak et al. studied
vocabulary changes in more than 15 million biomedical abstracts from 2010 to 2024 indexed by PubMed and show how the appearance of LLMs led to an abrupt increase in the frequency of certain style words. This excess word analysis suggests that at least 13.5% of 2024 abstracts were processed with LLMs.
Although they say that the analysis was performed on the corpus level and cannot identify individual texts that may have been processed by a LLM, we can of course check the proportion of LLM words in a text.
Unfortunately their online list contains stop words that I am eliminating here. But then we can run the following script!
# based on https://github.com/berenslab/llm-excess-vocab/tree/main
import csv
import re
import os
from collections import Counter
from striprtf.striprtf import rtf_to_text
from nltk.corpus import stopwords
import nltk
import chardet
# Ensure stopwords are available
nltk.download('stopwords')
# Paths
rtfd_folder_path = '/Users/x/Desktop/mss_image.rtfd' # RTFD is a directory
rtf_file_path = os.path.join(rtfd_folder_path, 'TXT.rtf') # or 'index.rtf'
csv_file_path = '/Users/x/Desktop/excess_words.csv'
# Read and decode the RTF file
with open(rtf_file_path, 'rb') as f:
raw_data = f.read()
# Try decoding automatically
encoding = chardet.detect(raw_data)['encoding']
rtf_content = raw_data.decode(encoding)
plain_text = rtf_to_text(rtf_content)
# Normalize and tokenize text
words_in_text = re.findall(r'\b\w+\b', plain_text.lower())
# Remove stopwords
stop_words = set(stopwords.words('english'))
filtered_words = [word for word in words_in_text if word not in stop_words]
# Load excess words from CSV
with open(csv_file_path, 'r', encoding='utf-8') as csv_file:
reader = csv.reader(csv_file)
excess_words = {row[0].strip().lower() for row in reader if row}
# Count excess words in filtered text
excess_word_counts = Counter(word for word in filtered_words if word in excess_words)
# Calculate proportion
total_words = len(filtered_words)
total_excess = sum(excess_word_counts.values())
proportion = total_excess / total_words if total_words > 0 else 0
# Output
print("\nExcess Words Found (Sorted by Frequency):")
for word, count in excess_word_counts.most_common():
print(f"{word}: {count}")
print(f"\nTotal words (without stopwords): {total_words}")
print(f"Total excess words: {total_excess}")
print(f"Proportion of excess words: {proportion:.4f}")
7 Aug 2025
The long ’em dash’ — U+2014 instead of the standard minus – seems to be a characteristic sign of chatGPT 4 even when asked not use it.
After giving up Nextcloud – which is now overkill for me with 30.000 files of basic setup – I am syncing now using iCloud. As I am working only on the desktop, it would make sense to sync the desktop in regular intervals but unfortunately this can be done only together with the Documents folder (something I don’t want). SE has also no good solution, so here is mine
# make another Desktop in iCloud folder
mkdir -p ~/Library/Mobile\ Documents/com~apple~CloudDocs/iCloudDesktop
# sync local Desktop
rsync -av --delete ~/Desktop/ ~/Library/Mobile\ Documents/com~apple~CloudDocs/iCloudDesktop/
# and run it every hour or so
# launchctl load ~/Library/LaunchAgents/launched.com.desktop.rsync.plist
<?xml version="1.0" encoding="UTF-8"?>
<!DOCTYPE plist PUBLIC "-//Apple//DTD PLIST 1.0//EN" "http://www.apple.com/DTDs/PropertyList-1.0.dtd">
<plist version="1.0">
<dict>
<key>KeepAlive</key>
<dict>
<key>Crashed</key>
<true/>
</dict>
<key>Label</key>
<string>launched.com.desktop.rsync</string>
<key>ProgramArguments</key>
<array>
<string>/usr/bin/rsync</string>
<string>-av</string>
<string>--delete</string>
<string>/Users/xxx/Desktop/</string>
<string>/Users/xxx/Library/Mobile Documents/com~apple~CloudDocs/iCloudDesktop/</string>
</array>
<key>RunAtLoad</key>
<true/>
<key>StartCalendarInterval</key>
<array>
<dict>
<key>Minute</key>
<integer>0</integer>
</dict>
</array>
<key>StandardOutPath</key>
<string>/tmp/rsync.out</string>
<key>StandardErrorPath</key>
<string>/tmp/rsync.err</string>
</dict>
</plist>
There is a funny paper at arXiv, that is now published in Neurology. It claims to have found a
neural conundrum behind the slowness of human behavior. The information throughput of a human being is about 10 bits/s. In comparison, our sensory systems gather data at ~10^9 bits/s. The stark contrast between these numbers remains unexplained and touches on fundamental aspects of brain function: What neural substrate sets this speed limit on the pace of our existence? Why does the brain need billions of neurons to process 10 bits/s? Why can we only think about one thing at a time?
If there are really two brains, an “outer” brain with fast high-dimensional sensory and motor signals and an “inner” brain that does are the processing? My inner brain says this is a huge speculation.
I needed this urgently for indexing PDFs as Spotlight on the Mac is highly erratic after all this years.
Anything LLM seemed the most promising approach with an easy to use GUI and being well documented. But indexing failed after several hours, so I went on with LM Studio. Also this installation turned out to be more complicated than expected due to library “dependency hell” and version mismatch spiralling…
30.000 PDFs result in a 4G index while the system is unfortunately not very responsive (yet)
Continue reading How to run LLaMA on your local PDFs
Here is our newest paper – a nice collaboration with Andrea Taloni et al. along with a nice commentary – to recognize surgisphere-like fraud
Recently, it was proved that the large language model Generative Pre-trained Transformer 4 (GPT-4; OpenAI) can fabricate synthetic medical datasets designed to support false scientific evidence. To uncover statistical patterns that may suggest fabrication in datasets produced by large language models and to improve these synthetic datasets by attempting to remove detectable marks of nonauthenticity, investigating the limits of generative artificial intelligence.
[…] synthetic datasets were produced for 3 fictional clinical studies designed to compare the outcomes of 2 alternative treatments for specific ocular diseases. Synthetic datasets were produced using the default GPT-4o model and a custom GPT. Data fabrication was conducted in November 2024. Prompts were submitted to GPT-4o to produce 12 “unrefined” datasets, which underwent forensic examination. Based on the outcomes of this analysis, the custom GPT Synthetic Data Creator was built with detailed instructions to generate 12 “refined” datasets designed to evade authenticity checks. Then, forensic analysis was repeated on these enhanced datasets. […]
Sufficiently sophisticated custom GPTs can perform complex statistical tasks and may be abused to fabricate synthetic datasets that can pass forensic analysis as authentic.
Lensrental has some great advice
Quantity Based: One of the continual problems the AI art generation faces is in quantity, though it is continually improving. For instance, in the past, AI art would struggle with getting the correct number of fingers correct, or perhaps the correct placement of knuckles and joints in the fingers.
General Softness & Low Resolution: AI art takes immense computing power to generate, and it still hasn’t streamlined this problem. So often, AI art is limited in resolution and detail.
Repetition: To further expand on the tip above, AI art often uses repetition to help speed up the generation process. So you may see something copied several times over the same image.
Asymmetry: Asymmetry exists in all facets of life, [… if you] photograph the building so that it looks symmetrical across the plane. AI doesn’t understand these rules and often creates subtle symmetry shifts in its images.
TBC

Although promised, the supplement with the questions is missing and there is no information in the methods how the authors prompted – so basically useless as methods cannot be replicated.
And at the end, chatGPT is only a mirror of how good or bad these experts informed the public…
I consult the almighty chatGPT frequently for additional information as this saves me hours of wading through my own database, Pubmed, Scholar and Goggle Hits.
But I have my own opinion, I never cut & paste as this is always running at risk (1) to plagiarize unknowingly and (2) to produce nonsense.
Miryam Naddaf has an article about this
In a survey of nearly 5,000 researchers, some 19% said they had already tried using LLMs to ‘increase the speed and ease’ of their review. But the survey, by publisher Wiley, headquartered in Hoboken, New Jersey, didn’t interrogate the balance between using LLMs to touch up prose, and relying on the AI to generate the review.
And well, maybe I am already sticking to the NEJM that said
Although human expert review should continue to be the foundation of the scientific process, LLM feedback could benefit researchers
There is a super interesting study on citation quality in scientific studies.
In this study, we introduced a computational pipeline to analyze citation fidelity on a large scale and developed an automated measure to evaluate how authors report prior findings. By applying this method to 13 million citation sentence pairs, we uncovered several key insights into the dynamics of scholarly communication and show citation fidelity is higher with closer proximity. Through a quasi-causal experiment, we further establish that the “telephone effect” contributes to information loss: when citing papers exhibit low fidelity to the original claim, future papers that cite both the citing paper and the original show even lower fidelity to the original.

Clearly incorrect or misleading information couldn’t be identified in this study although the definition a cut-off should be possible.
And well, I would be also very interested if there are differences by scientific field.