for living then longer … This might be a good rule for private life (and even for a scientific career) but not so much for the progress of science.
There is another allergy gene paper on FCER1A and RAD50. FCER1A has some tragedy as the authors believed for many years in FCER1B (and others in FCER2).
Another tragedy comes with the second gene – Continue reading Try to stay away from rants and comparisons
If a thought arises, take note of it and then dismiss it
Can we really control our thoughts? A PLoS ONE paper is looking into neural correlates of distraction by comparing 12 Zen meditators and 12 control subjects that were offered a “real English word” or “not a real English word” while resting in a 3 Tesla Siemens Magnetom. Outcome has been the time of refocusing attention to the breathing. The results are quite convincing (and not really unexpected) that the Zen meditators showed a reduced duration of their neural response to distraction probably by cutting down emotional self-reflectance and other associations coming up. I would be happy to participate also in a trial where somebody could record my EEG while driving on the of these new 185 kW Tesla roadsters.
ORMDL3 – more inconsistencies in the Nature paper
It seems that CEA just published my rather critical view of the recent ORMDL3 association paper. My letter had been first submitted to “Nature” but rejected after review.
A rebuttal seemed to be necessary as the authors repeatedly highlight ORMDL3 as a new asthma gene – in the printed Nature paper, in the Nature podcast and in the accompanying press releases. They even continue with reviews saying that
Completion of the human genome sequence and the advent of genome-wide association studies have resulted in the identification of two novel asthma susceptibility genes, ORMDL3 and CHI3L1, in the past year.
Continue reading ORMDL3 – more inconsistencies in the Nature paper
The largest experiment of the world
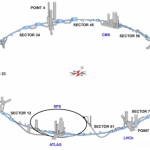 Physics gets now a lot of attention with the largest experiment ever done with the LHC (pictures+data).
Physics gets now a lot of attention with the largest experiment ever done with the LHC (pictures+data).
* largest machine (26.659 m diameter)
* largest freezer (60 tons helium)
* fastest runaway (99,99999 % speed of light)
* highest energy consumption (7 TeV)
* most lonely place – vaccum (10^-13 at)
* hottest place 100.000-fold temperature of the sun
* coldest place onearth (-271,3°C)
* largest supercomputer
Gratulations from the biomedical field that produced much less records in this week.
SNP batch annotation of GWAs
Genowatch (paper|website) is doing pretty well by annotating large SNP sets that would require otherwise numerous hours to map their position on genes, biological function and pathways. Continue reading SNP batch annotation of GWAs
Galaxy work flows
 Galaxy, one of my favorite bioinformatics websites now offers the conversion of existing history into a work flow. Obviously source data (by UCSC or Ensembl) may be produced in the same format but everything else can then be delegated to a workflow – just a set of instructions how to modify your data.
Galaxy, one of my favorite bioinformatics websites now offers the conversion of existing history into a work flow. Obviously source data (by UCSC or Ensembl) may be produced in the same format but everything else can then be delegated to a workflow – just a set of instructions how to modify your data.
The end of the hygiene hypothesis
The authors put a question mark at the end of the above statement while I would not hesitate to put an exclamation mark there. Writing this as a comment to a new study in the IJE they summarize the evidence that the “epidemics” of asthma in Western countries has begun to decline – as hygiene standards are not declining this might indicate the end of the hygiene hypothesis. Continue reading The end of the hygiene hypothesis
Dissent over descent
Having a free copy of the Lancet at the moment, I found a nice book review about “Dissent over Descent” by Steven Rose.
[He] takes a pleasure, which in part I share, in puncturing the often hyperbolic claims of natural scientists to be unimpeachable purveyors of absolute truth Continue reading Dissent over descent
The pointillism GWA plot
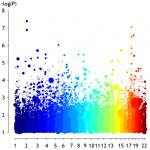
The experts in the field will immediately notice what I am suggesting here – an improved GWA plot that does not take into account p values alone but also effect sizes. I was experimenting some time with smile plots but finally ended with this bubble plot. Bubble size for 0.5<OR>2 is set to a minimum while all other ORs get increasing bubbles (BTW use for OR<1 a 1/OR transformation beforehand). Chromosomal colors are from a self defined palette using the colorRampPalette function in R which makes it look like pointillism art. The real question: Did the previous GWA p value screening miss some important effects? For example the important dot at x=4 and y=4?
Common disease + common variant = common misunderstanding?
Three large studies on schizophrenia now all agree that rare structural variants (CNVs) have a causal role in disease causation – probably by affecting large regions at 1q21 and 15q11 that contain all some “neuro”-genes (Harvard, Decode, Seattle). There are two major points that let me wonder if these papers even mark a turning point in our understanding of genetic causes of human diseases that is largely influenced by the Chakravarti hypothesis of common disease and common variants (CDVC). The Seattle paper had the most clever comment on that
We propose an alternative model: that some mutations predisposing to schizophrenia are highly penetrant, individually rare, and of recent origin, even specific to single cases or families.
Continue reading Common disease + common variant = common misunderstanding?
Continuous partial attention – a frequent scientist disease
Originally coined by Linda Stone 2007 as a Harvard Business Review “Breakthrough Idea” this gets a rapidly expanding disease among scientists and journal editors. A New Atlantis essay has some more details on the Myth of multitasking Continue reading Continuous partial attention – a frequent scientist disease
Fulchester graduates
Nature June, 19, 2008, has a nice correspondence: Fewer academics could be the answer to insufficient grants. A British author writes what many think but nobody wants to say. Read more about the country of Euphoria, his four universities, each with ten academics and the ambitious president of Fulchester … Unfortunately there is no proposal Continue reading Fulchester graduates
WikiPathways
 As somebody who is dealing most time with large datasets I always arrive at genes and proteins that I do not know. Using Biocarta, Keggs and other services in the past, I find the new WikiPathways exciting and hope that it will grow over the years. A companion paper in PLoS biology describes its roots in GenMAPP and the current work of the authors on bots that identifiy inconsistencies but also pick up loose ends. Hopefully I will find some time to work a bit on nuclear receptors, yea, yea.
As somebody who is dealing most time with large datasets I always arrive at genes and proteins that I do not know. Using Biocarta, Keggs and other services in the past, I find the new WikiPathways exciting and hope that it will grow over the years. A companion paper in PLoS biology describes its roots in GenMAPP and the current work of the authors on bots that identifiy inconsistencies but also pick up loose ends. Hopefully I will find some time to work a bit on nuclear receptors, yea, yea.
Dung hill counting
The logo site
Sequence logos are fun: www-lmmb.ncifcrf.gov is the ultimate site for many demo figures. It provides even a link where you can manufacture your own logo. Here is mine which is derived from a TCR consensus sequence: