Category Archives: Software
Vanish
Finally, here is a technical solution to a proposal that I made here earlier
Ok, we are aware that recovery is always possible with cut & paste into other applications or printing a text -so we may better think about some watermarked graphics. “Don’t ever say anything on e-mail or text messaging that you don’t want to come back and bite you.”
Do computer games lead to aggression?
We had a lunch discussion on that topic this week – of course, investigating the computer of a person running amok will reveal some (Counterstrike type) games due to his isolation. But such games are also on million of othercomputers who do not run riot and who get relaxed by playing games.
Nevertheless I read this week about the aggressive behaviour of teens who were not allowed to play the whole night World of Warcraft. So, there seems to be clear links Continue reading Do computer games lead to aggression?
P≤0.0069 in election results
Too many “7” digits were found for candidate A in a study of the presidential election results.
This method is closely related to Benford’s Law. A highly significant (p ~ 0.0007) excess of vote counts for candidate K that start with the digit 7 is found (41 observed, 21.2–22 expected).
I have to admit that I heard for the first time of the Newcomb-Benford’s Law (NBL)…
Please, send always an ics file in the attachment
as it is rather time consuming to type in all appointments.
Appointments can be exported from outlook (win XP) abd ical (Mac OS X) just by drag and drop to your favorite mail program. It is just one click here on the attachment to import the date here (Lufthansa already knows that but you may use this hint in your company to get the inventor award). BTW The screenshot shows another neat idea of integrating dynamic Doodle feeds.
Genome browser introduces error lane
A welcome initative of the Golden Path Genome Browser:
Users often send us notes and hints about how we might better curate and annotate the genome. These contributions are valuable but it isn’t feasible for us to update our annotations on a per-user basis. Continue reading Genome browser introduces error lane
If it wasn’t actually you
this is at least what Alexa tells me about you – what a nonsense giving averages of so different visitors ;-)
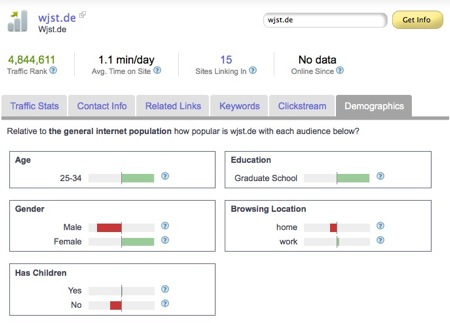
although the profile isn’t so bad at al l…
What is the best logistic model?
I have never heard a formal lecture answering this question even after many years in epidemiology. It should be parsimonious of course to avoid too many missings but seems largely a subjective approach to keep or drop a variable. It was therefore quite helpful to find now an online lecture that exemplifies a sound approach – check out unc.edu/courses/2006spring. I already used anova to compare models (at least since my move from SAS to R) while using AIC is something that I am adding now to my toolbox. Continue reading What is the best logistic model?
Mac OS X running Sigmaplot
 I haven’t found any replacement yet for Sigmaplot that is offered only as a native Windows application. Even buying a copy of Aabel didn’t bring me back the fast and intuitive plotting capabilities. Running Sigmaplot under Parallels is a pain using too much of the system ressources while Crossover failed to start up even after following these step-by-step instructions. Finally, with Crossover 8, it seems that a win98 bottle can run Sigmaplot 9!
I haven’t found any replacement yet for Sigmaplot that is offered only as a native Windows application. Even buying a copy of Aabel didn’t bring me back the fast and intuitive plotting capabilities. Running Sigmaplot under Parallels is a pain using too much of the system ressources while Crossover failed to start up even after following these step-by-step instructions. Finally, with Crossover 8, it seems that a win98 bottle can run Sigmaplot 9!
Science flies
Still in the spirit of the last few posts, here comes something exciting: sciflies.org aims at
We look forward to receiving your application for funding of initial proof-of-concept STEM research projects in the range of $5,000 to $12,000. To participate in this unique online grassroots-funded opportunity, please complete the questionnaire about your project, including details of its possible outcome/impact and profiles of the researchers or research team.
but, sorry, I have to warn you – the website does NOT save your project – it took me 20 minutes to figure that out.
10% of the energy consumption of computers by unwanted threads?
The title already says it all – I think that poorly programmed software leads a lot of traces on the harddisk, in the memory and even on the CPU load.
It took me more than 1 hour and even several emails to figure where launchd starts this process:
Finally, just silence, several months after deinstallation of the client, yea, yea.
Complains about the next generation
This seems to have a long tradition going even back to Plato’s republic. Sure – the demonstrated fierceness covers only the crudity of the growing identity. Continue reading Complains about the next generation
Mirror neurons and science careers
Spiegel online has an excellent report about mirror neurons, empathy, social background and research (taking up a theme in the ZEIT 2003)
“In unserer Kultur sind am erfolgreichsten die”, sagt Gruen, “die am meisten von ihren Gefühlen, von der Fähigkeit zum Mitgefühl abgeschnitten sind.”
Continue reading Mirror neurons and science careers
Why just C11orf30?
I have already seen this data in Washington and even talked to one of the two first authors (pun!) at the airport – the first genomewide scan for atopic dermatitis is now being online at the nature genetics website. The overall effects are disppointing small – my quick plot gives the cumulative (sic!) negative log p values.
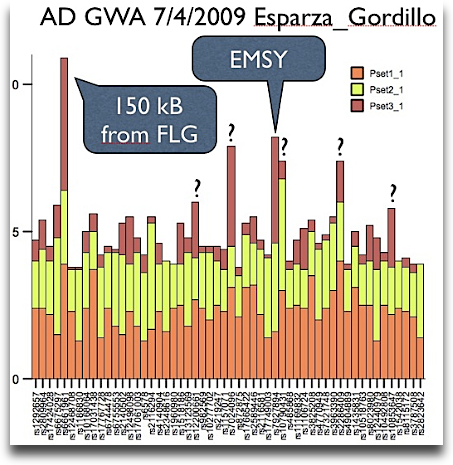
Continue reading Why just C11orf30?
I don’t want to bid for this dinner
NYT reports that Knome plans to offer its personal gene-sequencing service to the highest bidder in an eBay auction set to begin on Friday and continue for 10 days. This will include also a private dinner Continue reading I don’t want to bid for this dinner