Also outside the genetics community many people wonder why Popper’s account of falsifiability has so readily be abandoned. Karl Popper used falsification in “Logic of Scientific Discovery” as a criterion of demarcation between what is and what is not genuinely scientific.
Paul K. Feyerabend, one of Poppers many famous scholars at the London School of Economics- defended in “Against Method” (Feyerabend 1993) the view that there are no methodological rules which can be always used by scientists. He objected to any single prescriptive scientific method (like falsification) as any such method would limit the activities of scientists, and restrict scientific progress. Progress instead occurs where new theories are not consistent with older theories; a new theory also can never be consistent with all relevant facts: this make falsification attempts useless. Feyerabend advocated in a rather anarchistic view that scientific pluralism improves the critical power of science and not any schematic rules like profile population x with SNP panel y and describe all p less than z to finally develop new treatment t.
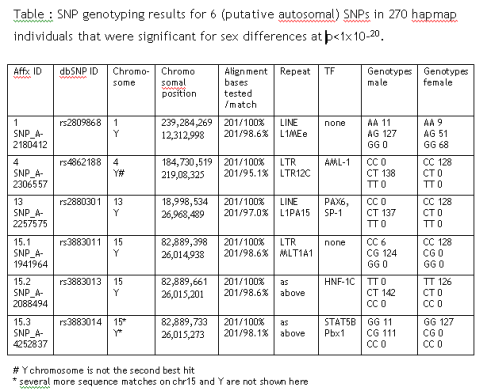
Many reasons why genetic association studies failed have been already identified (see Buchanan et al. 2006). Usually high impact journals get spectacular claims first; half-way down between Popper and Feyerabend, the editorial board looks for falsifiability by claiming additional populations.
As expected, effect sizes will not be exactly the same in different populations; often only neighbouring SNP “rescue” the initial claim. It has never been decided by a formal process, what does it mean if a third or fourth population doesn’t show up with the same result. It has never been clarified that falsifiability means that the exactly same SNP needs to be associated in all population studies or just a haplotype (or just a microsatellite allele) somewhere in that genomic region.
Nevertheless replication validity – in the context of generalization – is permanently used to prove “true” or “meaningful” association that ultimately deserve a high impact factor. Humans look different and they seem different in genetic terms: the high individual variablity in expressing a disease trait may reflect not only reflect a highly variable environment but also highly individual genetic pathway. We are willing to accept a causal mutation found in just one family with a monogenic trait often there seems no way to convince an editorial board that a strong association found in just one study sample is an important discovery that may severely impact exactly this population (given additional functional proof of otherwise static gene variant).
The absence of large linkage signals and the absence of reproducible genetic associations with nearly all complex diseases may indicate only individual risk gene combinations. It seems to be that we need to listen to another scholar of Popper – Thomas Kuhn — to change the current paradigma.
Addendum
14-6-07 Finally, Nature published some guidelines for interpretation of association studies
CC-BY-NC Science Surf , accessed 06.06.2026